CAS10-BASED ASSAY FOR NUCLEIC ACID DETECTION
US20240191217
2024-06-13
Chemistry; metallurgy
C12N9/22
Inventors:
Applicant:
Drawings (4 of 15)
Smart overview of the Invention
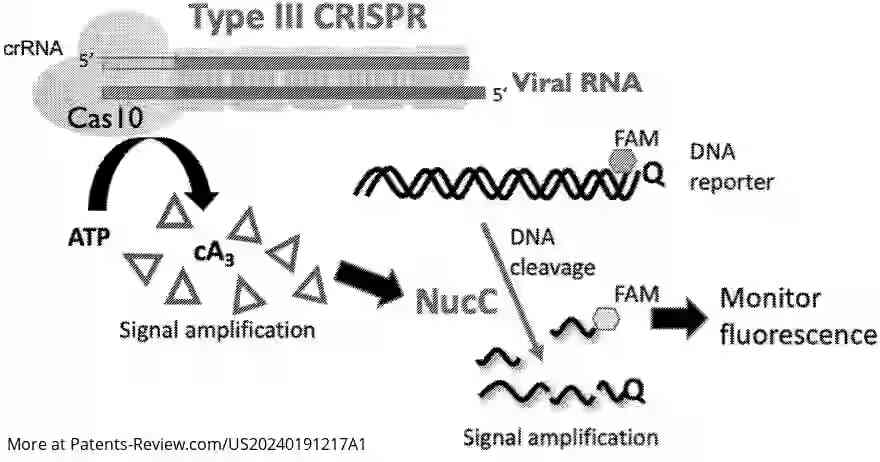
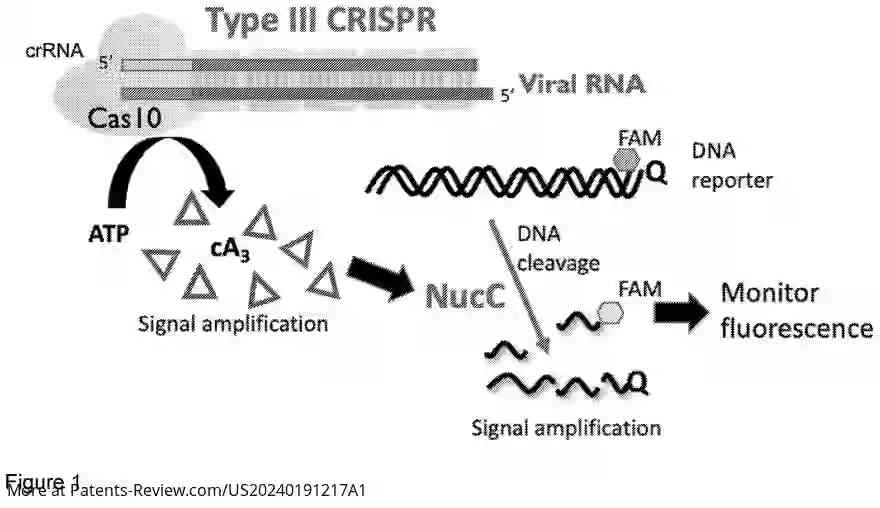
The patent application describes a novel detection system for nucleic acids using a CRISPR-associated effector protein, Cas10. This system is designed to identify target nucleic acids by complementary base pairing, leading to the production of cyclic oligoadenylate (cOA) upon RNA binding. The cOA activates a nuclease, such as NucC, which through a reporter system, produces a detectable fluorescent or colored signal. The preferred embodiments utilize Cas10 and NucC from Vibrio metoccus.
Background and Need
Infectious diseases like Covid-19 pose significant public health challenges, necessitating reliable detection methods. Current gold standard tests, such as RT-PCR, require complex equipment and are time-consuming. Alternative methods like DETECTR and SHERLOCK use CRISPR effectors but still involve DNA conversion steps. The disclosed method aims to provide a faster, simpler CRISPR-based approach for detecting infectious agents directly from samples.
Methodology
The proposed system uses Cas10 to detect specific RNA sequences and generate cOA, which activates a nuclease to produce a detectable signal. This method can be tailored for various target nucleic acids by adjusting the molecule that binds the target sequence. The advantage lies in its ability to produce substantial amounts of cOA per detected RNA molecule, enhancing detection sensitivity.
Applications
This detection system can be applied in diagnosing viral pathogens and related diseases. It is particularly useful for detecting RNA from viruses such as SARS-COV-2, SARS-COV-1, and MERS-COV. By identifying viral RNA sequences, it aids in diagnosing conditions like Covid-19, SARS, and MERS.
Advantages and Specificity
A key benefit of this system is its low limit of detection (LOD), capable of identifying very low concentrations of target nucleic acids without needing amplification steps. For instance, it can achieve LODs as low as <1 fM for viral nucleic acids like those from SARS-COV-2. This high sensitivity makes it suitable for detecting specific genes or transcripts, including those from pathogens or viruses.